cvshrink
Cross-validate regularization of linear discriminant
Syntax
Description
[___] = cvshrink(
specifies additional options using one or more name-value arguments. For example, you can
specify the number of delta and gamma intervals for cross-validation, and the verbosity
level of progress messages. Mdl,Name=Value)
Examples
Input Arguments
Name-Value Arguments
Output Arguments
More About
Tips
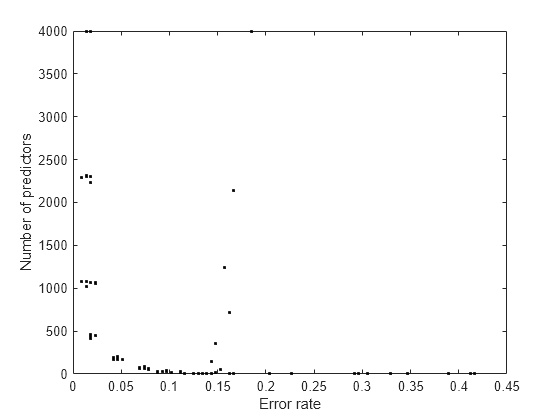
Examine the
errandnumpredoutputs to see the tradeoff between the cross-validated error and the number of predictors. When you find a satisfactory point, set the correspondinggammaanddeltaproperties in the model using dot notation. For example, if(i,j)is the location of the satisfactory point, set:Mdl.Gamma = gamma(i); Mdl.Delta = delta(i,j);
Version History
Introduced in R2012b