margin
Classification margins for multiclass error-correcting output codes (ECOC) model
Syntax
Description
m = margin(Mdl,tbl,ResponseVarName)m) for the trained multiclass error-correcting output codes (ECOC)
model Mdl using the predictor data in table tbl
and the class labels in tbl.ResponseVarName.
m = margin(___,Name,Value)
Examples
Calculate the test-sample classification margins of an ECOC model with SVM binary learners.
Load Fisher's iris data set. Specify the predictor data X, the response data Y, and the order of the classes in Y.
load fisheriris X = meas; Y = categorical(species); classOrder = unique(Y); % Class order rng(1) % For reproducibility
Train an ECOC model using SVM binary classifiers. Specify a 30% holdout sample, standardize the predictors using an SVM template, and specify the class order.
t = templateSVM('Standardize',true); PMdl = fitcecoc(X,Y,'Holdout',0.30,'Learners',t,'ClassNames',classOrder); Mdl = PMdl.Trained{1}; % Extract trained, compact classifier
PMdl is a ClassificationPartitionedECOC model. It has the property Trained, a 1-by-1 cell array containing the CompactClassificationECOC model that the software trained using the training set.
Calculate the test-sample classification margins. Display the distribution of the margins using a boxplot.
testInds = test(PMdl.Partition); % Extract the test indices XTest = X(testInds,:); YTest = Y(testInds,:); m = margin(Mdl,XTest,YTest); boxplot(m) title('Test-Sample Margins')
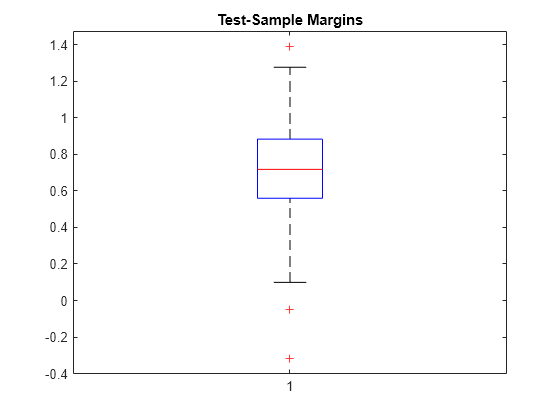
The classification margin of an observation is the positive-class negated loss minus the maximum negative-class negated loss. Choose classifiers that yield relatively large margins.
Perform feature selection by comparing test-sample margins from multiple models. Based solely on this comparison, the model with the greatest margins is the best model.
Load Fisher's iris data set. Specify the predictor data X, the response data Y, and the order of the classes in Y.
load fisheriris X = meas; Y = categorical(species); classOrder = unique(Y); % Class order rng(1); % For reproducibility
Partition the data set into training and test sets. Specify a 30% holdout sample for testing.
Partition = cvpartition(Y,'Holdout',0.30); testInds = test(Partition); % Indices for the test set XTest = X(testInds,:); YTest = Y(testInds,:);
Partition defines the data set partition.
Define these two data sets:
fullXcontains all four predictors.partXcontains the sepal measurements only.
fullX = X; partX = X(:,1:2);
Train an ECOC model using SVM binary classifiers for each predictor set. Specify the partition definition, standardize the predictors using an SVM template, and define the class order.
t = templateSVM('Standardize',true); fullPMdl = fitcecoc(fullX,Y,'CVPartition',Partition,'Learners',t,... 'ClassNames',classOrder); partPMdl = fitcecoc(partX,Y,'CVPartition',Partition,'Learners',t,... 'ClassNames',classOrder); fullMdl = fullPMdl.Trained{1}; partMdl = partPMdl.Trained{1};
fullPMdl and partPMdl are ClassificationPartitionedECOC models. Each model has the property Trained, a 1-by-1 cell array containing the CompactClassificationECOC model that the software trained using the corresponding training set.
Calculate the test-sample margins for each classifier. For each model, display the distribution of the margins using a boxplot.
fullMargins = margin(fullMdl,XTest,YTest); partMargins = margin(partMdl,XTest(:,1:2),YTest); boxplot([fullMargins partMargins],'Labels',{'All Predictors','Two Predictors'}) title('Boxplots of Test-Sample Margins')
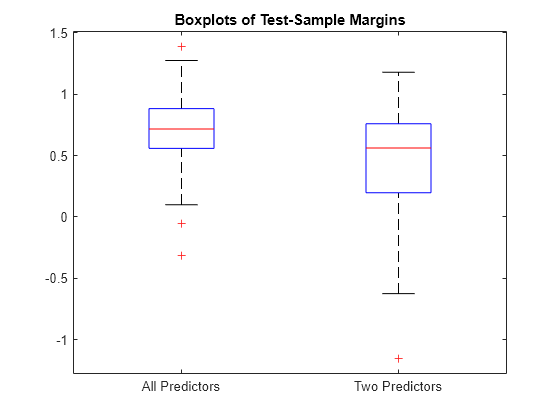
The margin distribution of fullMdl is situated higher and has less variability than the margin distribution of partMdl.
Input Arguments
Full or compact multiclass ECOC model, specified as a
ClassificationECOC or
CompactClassificationECOC model
object.
To create a full or compact ECOC model, see ClassificationECOC or CompactClassificationECOC.
Sample data, specified as a table. Each row of tbl corresponds to one
observation, and each column corresponds to one predictor variable. Optionally,
tbl can contain additional columns for the response variable
and observation weights. tbl must contain all the predictors used
to train Mdl. Multicolumn variables and cell arrays other than cell
arrays of character vectors are not allowed.
If you train Mdl using sample data contained in a
table, then the input data for margin
must also be in a table.
When training Mdl, assume that you set
'Standardize',true for a template object specified in the
'Learners' name-value pair argument of fitcecoc. In
this case, for the corresponding binary learner j, the software standardizes
the columns of the new predictor data using the corresponding means in
Mdl.BinaryLearner{j}.Mu and standard deviations in
Mdl.BinaryLearner{j}.Sigma.
Data Types: table
Response variable name, specified as the name of a variable in tbl. If
tbl contains the response variable used to train
Mdl, then you do not need to specify
ResponseVarName.
If you specify ResponseVarName, then you must do so as a character vector
or string scalar. For example, if the response variable is stored as
tbl.y, then specify ResponseVarName as
'y'. Otherwise, the software treats all columns of
tbl, including tbl.y, as predictors.
The response variable must be a categorical, character, or string array, a logical or numeric vector, or a cell array of character vectors. If the response variable is a character array, then each element must correspond to one row of the array.
Data Types: char | string
Predictor data, specified as a numeric matrix.
Each row of X corresponds to one observation, and each column corresponds
to one variable. The variables in the columns of
X must be the same as the
variables that trained the classifier
Mdl.
The number of rows in X must equal the number of rows in
Y.
When training Mdl, assume that you set
'Standardize',true for a template object specified in the
'Learners' name-value pair argument of fitcecoc. In
this case, for the corresponding binary learner j, the software standardizes
the columns of the new predictor data using the corresponding means in
Mdl.BinaryLearner{j}.Mu and standard deviations in
Mdl.BinaryLearner{j}.Sigma.
Data Types: double | single
Class labels, specified as a categorical, character, or string array, a logical or numeric
vector, or a cell array of character vectors. Y must have the same
data type as Mdl.ClassNames. (The software treats string arrays as cell arrays of character
vectors.)
The number of rows in Y must equal the number of rows in
tbl or X.
Data Types: categorical | char | string | logical | single | double | cell
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: margin(Mdl,tbl,'y','BinaryLoss','exponential') specifies an
exponential binary learner loss function.
Binary learner loss function, specified as a built-in loss function name or function handle.
This table describes the built-in functions, where yj is the class label for a particular binary learner (in the set {–1,1,0}), sj is the score for observation j, and g(yj,sj) is the binary loss formula.
Value Description Score Domain g(yj,sj) "binodeviance"Binomial deviance (–∞,∞) log[1 + exp(–2yjsj)]/[2log(2)] "exponential"Exponential (–∞,∞) exp(–yjsj)/2 "hamming"Hamming [0,1] or (–∞,∞) [1 – sign(yjsj)]/2 "hinge"Hinge (–∞,∞) max(0,1 – yjsj)/2 "linear"Linear (–∞,∞) (1 – yjsj)/2 "logit"Logistic (–∞,∞) log[1 + exp(–yjsj)]/[2log(2)] "quadratic"Quadratic [0,1] [1 – yj(2sj – 1)]2/2 The software normalizes binary losses so that the loss is 0.5 when yj = 0. Also, the software calculates the mean binary loss for each class [1].
For a custom binary loss function, for example
customFunction, specify its function handleBinaryLoss=@customFunction.customFunctionhas this form:bLoss = customFunction(M,s)
Mis the K-by-B coding matrix stored inMdl.CodingMatrix.sis the 1-by-B row vector of classification scores.bLossis the classification loss. This scalar aggregates the binary losses for every learner in a particular class. For example, you can use the mean binary loss to aggregate the loss over the learners for each class.K is the number of classes.
B is the number of binary learners.
For an example of passing a custom binary loss function, see Predict Test-Sample Labels of ECOC Model Using Custom Binary Loss Function.
This table identifies the default BinaryLoss value, which depends on the
score ranges returned by the binary learners.
| Assumption | Default Value |
|---|---|
All binary learners are any of the following:
| "quadratic" |
| All binary learners are SVMs or linear or kernel classification models of SVM learners. | "hinge" |
All binary learners are ensembles trained by
AdaboostM1 or
GentleBoost. | "exponential" |
All binary learners are ensembles trained by
LogitBoost. | "binodeviance" |
You specify to predict class posterior probabilities by setting
FitPosterior=true in fitcecoc. | "quadratic" |
| Binary learners are heterogeneous and use different loss functions. | "hamming" |
To check the default value, use dot notation to display the BinaryLoss property of the trained model at the command line.
Example: BinaryLoss="binodeviance"
Data Types: char | string | function_handle
Decoding scheme that aggregates the binary losses, specified as
"lossweighted" or "lossbased". For more
information, see Binary Loss.
Example: Decoding="lossbased"
Data Types: char | string
Predictor data observation dimension, specified as the comma-separated pair consisting of
'ObservationsIn' and 'columns' or
'rows'. Mdl.BinaryLearners must contain
ClassificationLinear models.
Note
If you orient your predictor matrix so that
observations correspond to columns and specify
'ObservationsIn','columns', you
can experience a significant reduction in
execution time. You cannot specify
'ObservationsIn','columns' for
predictor data in a table.
Estimation options, specified as a structure array as returned by statset.
To invoke parallel computing you need a Parallel Computing Toolbox™ license.
Example: Options=statset(UseParallel=true)
Data Types: struct
Verbosity level, specified as 0 or 1.
Verbose controls the number of diagnostic messages that the
software displays in the Command Window.
If Verbose is 0, then the software does not display
diagnostic messages. Otherwise, the software displays diagnostic messages.
Example: Verbose=1
Data Types: single | double
Output Arguments
Classification margins, returned as a numeric column vector or numeric matrix.
If Mdl.BinaryLearners contains ClassificationLinear models, then m is an
n-by-L vector, where n is the
number of observations in X and L is the number
of regularization strengths in the linear classification models
(numel(Mdl.BinaryLearners{1}.Lambda)). The value
m(i,j) is the margin of observation i for the
model trained using regularization strength
Mdl.BinaryLearners{1}.Lambda(j).
Otherwise, m is a column vector of length
n.
More About
The binary loss is a function of the class and classification score that determines how well a binary learner classifies an observation into the class. The decoding scheme of an ECOC model specifies how the software aggregates the binary losses and determines the predicted class for each observation.
Assume the following:
mkj is element (k,j) of the coding design matrix M—that is, the code corresponding to class k of binary learner j. M is a K-by-B matrix, where K is the number of classes, and B is the number of binary learners.
sj is the score of binary learner j for an observation.
g is the binary loss function.
is the predicted class for the observation.
The software supports two decoding schemes:
Loss-based decoding [2] (
Decodingis"lossbased") — The predicted class of an observation corresponds to the class that produces the minimum average of the binary losses over all binary learners.Loss-weighted decoding [3] (
Decodingis"lossweighted") — The predicted class of an observation corresponds to the class that produces the minimum average of the binary losses over the binary learners for the corresponding class.The denominator corresponds to the number of binary learners for class k. [1] suggests that loss-weighted decoding improves classification accuracy by keeping loss values for all classes in the same dynamic range.
The predict, resubPredict, and
kfoldPredict functions return the negated value of the objective
function of argmin as the second output argument
(NegLoss) for each observation and class.
This table summarizes the supported binary loss functions, where yj is a class label for a particular binary learner (in the set {–1,1,0}), sj is the score for observation j, and g(yj,sj) is the binary loss function.
| Value | Description | Score Domain | g(yj,sj) |
|---|---|---|---|
"binodeviance" | Binomial deviance | (–∞,∞) | log[1 + exp(–2yjsj)]/[2log(2)] |
"exponential" | Exponential | (–∞,∞) | exp(–yjsj)/2 |
"hamming" | Hamming | [0,1] or (–∞,∞) | [1 – sign(yjsj)]/2 |
"hinge" | Hinge | (–∞,∞) | max(0,1 – yjsj)/2 |
"linear" | Linear | (–∞,∞) | (1 – yjsj)/2 |
"logit" | Logistic | (–∞,∞) | log[1 + exp(–yjsj)]/[2log(2)] |
"quadratic" | Quadratic | [0,1] | [1 – yj(2sj – 1)]2/2 |
The software normalizes binary losses so that the loss is 0.5 when yj = 0, and aggregates using the average of the binary learners [1].
Do not confuse the binary loss with the overall classification loss (specified by the
LossFun name-value argument of the loss and
predict object functions), which measures how well an ECOC classifier
performs as a whole.
The classification margin is, for each observation, the difference between the negative loss for the true class and the maximal negative loss among the false classes. If the margins are on the same scale, then they serve as a classification confidence measure. Among multiple classifiers, those that yield greater margins are better.
Tips
To compare the margins or edges of several ECOC classifiers, use template objects to specify a common score transform function among the classifiers during training.
References
[1] Allwein, E., R. Schapire, and Y. Singer. “Reducing multiclass to binary: A unifying approach for margin classifiers.” Journal of Machine Learning Research. Vol. 1, 2000, pp. 113–141.
[2] Escalera, S., O. Pujol, and P. Radeva. “Separability of ternary codes for sparse designs of error-correcting output codes.” Pattern Recog. Lett. Vol. 30, Issue 3, 2009, pp. 285–297.
[3] Escalera, S., O. Pujol, and P. Radeva. “On the decoding process in ternary error-correcting output codes.” IEEE Transactions on Pattern Analysis and Machine Intelligence. Vol. 32, Issue 7, 2010, pp. 120–134.
Extended Capabilities
The
margin function supports tall arrays with the following usage
notes and limitations:
margindoes not support talltabledata whenMdlcontains kernel or linear binary learners.
For more information, see Tall Arrays.
To run in parallel, specify the Options name-value argument in the call to
this function and set the UseParallel field of the
options structure to true using
statset:
Options=statset(UseParallel=true)
For more information about parallel computing, see Run MATLAB Functions with Automatic Parallel Support (Parallel Computing Toolbox).
Usage notes and limitations:
The
marginfunction does not support models trained using:Decision tree learners with surrogate splits
SVM learners for one-class classification
KNN learners that use the Kd-tree nearest neighbor search method, function handle distance metrics, or tie inclusion
For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2014b
See Also
ClassificationECOC | CompactClassificationECOC | edge | resubMargin | predict | fitcecoc | loss
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleccione un país/idioma
Seleccione un país/idioma para obtener contenido traducido, si está disponible, y ver eventos y ofertas de productos y servicios locales. Según su ubicación geográfica, recomendamos que seleccione: .
También puede seleccionar uno de estos países/idiomas:
Cómo obtener el mejor rendimiento
Seleccione China (en idioma chino o inglés) para obtener el mejor rendimiento. Los sitios web de otros países no están optimizados para ser accedidos desde su ubicación geográfica.
América
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)