glmval
Generalized linear model values
Syntax
Description
[
also computes 95% confidence bounds for the predicted values. ypred,yLo,yUp] = glmval(b,Xnew,Link,stats)stats is a
structure returned by the glmfit function. yLo and
yUp define a lower confidence bound of ypred-yLo,
and an upper confidence bound of ypred+yUp. Confidence bounds are
nonsimultaneous by default, and apply to the fitted curve, not to a new observation.
[___] = glmval(___,Name=Value) specifies
options using one or more name-value arguments in addition to any of the input argument
combinations in previous syntaxes. For example, you can specify the confidence level for the
confidence bounds.
Examples
Generate sample data using Poisson random numbers with two underlying predictors X(:,1) and X(:,2).
rng(0,"twister") % For reproducibility rndvars = randn(100,2); X = [2 + rndvars(:,1),rndvars(:,2)]; mu = exp(1 + X*[1;2]); y = poissrnd(mu);
Fit a generalized linear regression model to the sample data.
b = glmfit(X,y,"poisson");b is a 3-by-1 vector of regression coefficients.
Create data points for prediction.
[Xtest1,Xtest2] = meshgrid(-1:.5:3,-2:.5:2); Xnew = [Xtest1(:),Xtest2(:)];
Predict the responses at the data points using the fitted model. Use the log link function, which is appropriate for a Poisson distribution.
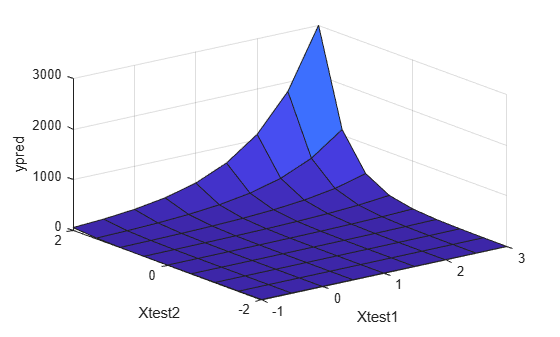
ypred = glmval(b,Xnew,"log");Plot the predictions.
surf(Xtest1,Xtest2,reshape(ypred,9,9)) xlabel("Xtest1") ylabel("Xtest2") zlabel("ypred")
Fit a generalized linear regression model, and compute predicted (estimated) values for the predictor data using the fitted model.
Create a sample data set.
x = [2100 2300 2500 2700 2900 3100 ...
3300 3500 3700 3900 4100 4300]';
n = [48 42 31 34 31 21 23 23 21 16 17 21]';
y = [1 2 0 3 8 8 14 17 19 15 17 21]';x contains the predictor variable values. Each y value is the number of successes in the corresponding number of trials in n.
Fit a probit regression model for y on x using a binomial distribution for y.
b = glmfit(x,[y n],"binomial",Link="probit");
b is a 2-by-1 vector of regression coefficients.
Compute the estimated number of successes.
ypred = glmval(b,x,"probit",BinomialSize=n);Plot the observed and estimated percent success fractions versus the x values.
plot(x,y./n,"o",x,ypred./n,"-") xlabel("x") ylabel("Success Fraction")

Create a sample data set.
x = [2100 2300 2500 2700 2900 3100 ...
3300 3500 3700 3900 4100 4300]';
n = [48 42 31 34 31 21 23 23 21 16 17 21]';
y = [1 2 0 3 8 8 14 17 19 15 17 21]';x contains the predictor variable values. Each y value is the number of successes in the corresponding number of trials in n.
Define three function handles, created by using @, that define the link, the derivative of the link, and the inverse link for a probit link function. Store the function handles in a structure.
link = @(mu) norminv(mu); derlink = @(mu) 1 ./ normpdf(norminv(mu)); invlink = @(resp) normcdf(resp); F = struct(Link=link,Derivative=derlink,Inverse=invlink);
Fit a generalized linear model for y on x using a binomial distribution for y and the link function that you defined.
b = glmfit(x,[y n],"binomial",F);b is a 2-by-1 vector of regression coefficients.
Compute the estimated number of successes.
ypred = glmval(b,x,F,BinomialSize=n);
Plot the observed and estimated percent success fractions versus the x values.
plot(x, y./n,"o",x,ypred./n,"-") xlabel("x") ylabel("Success fraction")

Train a generalized linear model, and then generate code from a function that classifies new observations based on the model. This example is based on the Use Custom-Defined Link Function example.
Enter the sample data.
x = [2100 2300 2500 2700 2900 3100 ...
3300 3500 3700 3900 4100 4300]';
n = [48 42 31 34 31 21 23 23 21 16 17 21]';
y = [1 2 0 3 8 8 14 17 19 15 17 21]';
Suppose that the inverse normal pdf is an appropriate link function for the problem.
Define a function named myInvNorm.m that accepts values of  and returns corresponding values of the inverse of the standard normal cdf.
and returns corresponding values of the inverse of the standard normal cdf.
function in = myInvNorm(mu) %#codegen %myInvNorm Inverse of standard normal cdf for code generation % myInvNorm is a GLM link function that accepts a numeric vector mu, and % returns in, which is a numeric vector of corresponding values of the % inverse of the standard normal cdf. % in = norminv(mu); end
Define another function named myDInvNorm.m that accepts values of  and returns corresponding values of the derivative of the link function.
and returns corresponding values of the derivative of the link function.
function din = myDInvNorm(mu) %#codegen %myDInvNorm Derivative of inverse of standard normal cdf for code %generation % myDInvNorm corresponds to the derivative of the GLM link function % myInvNorm. myDInvNorm accepts a numeric vector mu, and returns din, % which is a numeric vector of corresponding derivatives of the inverse % of the standard normal cdf. % din = 1./normpdf(norminv(mu)); end
Define another function named myInvInvNorm.m that accepts values of  and returns corresponding values of the inverse of the link function.
and returns corresponding values of the inverse of the link function.
function iin = myInvInvNorm(mu) %#codegen %myInvInvNorm Standard normal cdf for code generation % myInvInvNorm is the inverse of the GLM link function myInvNorm. % myInvInvNorm accepts a numeric vector mu, and returns iin, which is a % numeric vector of corresponding values of the standard normal cdf. % iin = normcdf(mu); end
Create a structure array that specifies each of the link functions. Specifically, the structure array contains fields named 'Link', 'Derivative', and 'Inverse'. The corresponding values are the names of the functions.
link = struct('Link','myInvNorm','Derivative','myDInvNorm',... 'Inverse','myInvInvNorm')
link =
struct with fields:
Link: 'myInvNorm'
Derivative: 'myDInvNorm'
Inverse: 'myInvInvNorm'
Fit a GLM for y on x using the link function link. Return the structure array of statistics.
[b,~,stats] = glmfit(x,[y n],'binomial','link',link);
b is a 2-by-1 vector of regression coefficients.
In your current working folder, define a function called classifyGLM.m that:
Accepts measurements with columns corresponding to those in
x, regression coefficients whose dimensions correspond tob, a link function, the structure of GLM statistics, and any validglmvalname-value pair argumentReturns predictions and confidence interval margins of error
function [yhat,lo,hi] = classifyGLM(b,x,link,varargin) %#codegen %CLASSIFYGLM Classify measurements using GLM model % CLASSIFYGLM classifies the n observations in the n-by-1 vector x using % the GLM model with regression coefficients b and link function link, % and then returns the n-by-1 vector of predicted values in yhat. % CLASSIFYGLM also returns margins of error for the predictions using % additional information in the GLM statistics structure stats. narginchk(3,Inf); if(isstruct(varargin{1})) stats = varargin{1}; [yhat,lo,hi] = glmval(b,x,link,stats,varargin{2:end}); else yhat = glmval(b,x,link,varargin{:}); end end
Generate a MEX function from classifyGLM.m. Because C uses static typing, codegen must determine the properties of all variables in MATLAB® files at compile time. To ensure that the MEX function can use the same inputs, use the -args argument to specify the following in the order given:
Regression coefficients
bas a compile-time constantIn-sample observations
xLink function as a compile-time constant
Resulting GLM statistics as a compile-time constant
Name
'Confidence'as a compile-time constantConfidence level 0.9
To designate arguments as compile-time constants, use coder.Constant.
codegen -config:mex classifyGLM -args {coder.Constant(b),x,coder.Constant(link),coder.Constant(stats),coder.Constant('Confidence'),0.9}
Code generation successful.
codegen generates the MEX file classifyGLM_mex.mexw64 in your current folder. The file extension depends on your system platform.
Compare predictions by using glmval and classifyGLM_mex. Specify name-value pair arguments in the same order as in the -args argument in the call to codegen.
[yhat1,melo1,mehi1] = glmval(b,x,link,stats,'Confidence',0.9); [yhat2,melo2,mehi2] = classifyGLM_mex(b,x,link,stats,'Confidence',0.9); comp1 = (yhat1 - yhat2)'*(yhat1 - yhat2); agree1 = comp1 < eps comp2 = (melo1 - melo2)'*(melo1 - melo2); agree2 = comp2 < eps comp3 = (mehi1 - mehi2)'*(mehi1 - mehi2); agree3 = comp3 < eps
agree1 = logical 1 agree2 = logical 1 agree3 = logical 1
The generated MEX function produces the same predictions as predict.
Input Arguments
Coefficient estimates, specified as a numeric column vector returned by the
glmfit function.
Data Types: single | double
New predictor input values, specified as a numeric matrix. Each row of
Xnew corresponds to one observation, and each column corresponds
to one predictor variable.
By default, glmval includes a constant term in the model.
Do not add a column of 1s directly to Xnew. You can change the
default behavior of glmval by specifying the
Constant name-value argument.
Data Types: single | double
Link function, specified as one of the following values.
| Link Function Name | Link Function | Mean (Inverse) Function |
|---|---|---|
"identity" | f(μ) = μ | μ = Xb |
"log" | f(μ) = log(μ) | μ = exp(Xb) |
"logit" | f(μ) = log(μ/(1–μ)) | μ = exp(Xb) / (1 + exp(Xb)) |
"probit" | f(μ) = Φ–1(μ), where Φ is the cumulative distribution function of the standard normal distribution. | μ = Φ(Xb) |
"comploglog" | f(μ) = log(–log(1 – μ)) | μ = 1 – exp(–exp(Xb)) |
"reciprocal" | f(μ) = 1/μ | μ = 1/(Xb) |
p (a number) | f(μ) = μp | μ = Xb1/p |
| f(μ) =
S.Link(μ) | μ =
S.Inverse(Xb) |
The link function defines the relationship f(μ) = Xnew*b between the mean response μ and the linear combination of predictors Xnew*b.
Data Types: char | string | single | double | struct
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: glmval(b,X,Link,Confidence=0.99) specifies a confidence level
of 99% for the confidence bounds.
Binomial size, specified as a numeric scalar which represents the number of trials
in a binomial model. You can also specify BinomialSize as a
numeric column vector with one value for each row of Xnew.
Example: BinomialSize=[10 20 10 40]'
Data Types: single | double
Confidence level of the confidence bounds for ypred,
specified as a numeric value from 0 to 1. The default value of 0.95 corresponds to the
95% confidence level.
Example: Confidence=0.99
Data Types: single | double
Flag to include a constant term (intercept) in the model, specified as either
"on" or "off".
"on"(default) —glmvalincludes a constant term in the model. The coefficient of the constant term is the first element ofb."off"—glmvalomits the constant term in the model.
Example: Constant="off"
Data Types: char | string
Offset variable in the model, specified as a numeric column vector with one value
for each row of Xnew.
glmval uses Offset as an additional
predictor with a coefficient value fixed at 1. In other words, the model formula is
f(μ) = Offset +
,XNew*b
where f is the link function,
μ is the mean response, and
XNew*b is the linear combination of
predictors XNew.
Example: Offset=ones(50,1)
Data Types: single | double
Flag to compute simultaneous confidence bounds, specified as a numeric or logical
1 (true) or 0
(false).
true—glmvalcalculates confidence bounds for the curve of response values corresponding to all predictor values inXnew, using Schefféʼs method. The range between the upper and lower bounds contains the curve that consists of true response values with 100*Confidence% confidence.false—glmvalcalculates confidence bounds for the response value at each observation inXnew. The confidence interval for a response value at a specific predictor value contains the true response value with 100*Confidence% confidence.
With simultaneous bounds, the entire curve of true response values is within the bounds at high confidence. By contrast, nonsimultaneous bounds require only the response value at a single predictor value to be within the bounds at high confidence. Therefore, simultaneous bounds are wider than nonsimultaneous bounds.
Example: Simultaneous=true
Data Types: char | string | logical
Output Arguments
Predicted generalized linear model values, returned as a numeric column vector with
the same number of rows as Xnew.
References
[1] Dobson, A. J. An Introduction to Generalized Linear Models. New York: Chapman & Hall, 1990.
[2] McCullagh, P., and J. A. Nelder. Generalized Linear Models. New York: Chapman & Hall, 1990.
[3] Collett, D. Modeling Binary Data. New York: Chapman & Hall, 2002.
Extended Capabilities
Usage notes and limitations:
| Argument | Notes and Limitations |
|---|---|
b |
|
Xnew | Must be a double- or single-precision numeric matrix |
Link |
|
stats | Must be a compile-time constant |
Constant name-value argument | Must be a compile-time constant |
| Name-value arguments | Names in name-value arguments must be compile-time constants. For
example, to specify a confidence level of 0.9, include
|
For more information on code generation, see Introduction to Code Generation for Statistics and Machine Learning Functions and Overview of Code Generation Using MATLAB Coder (MATLAB Coder).
This function fully supports GPU arrays. For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced before R2006a
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleccione un país/idioma
Seleccione un país/idioma para obtener contenido traducido, si está disponible, y ver eventos y ofertas de productos y servicios locales. Según su ubicación geográfica, recomendamos que seleccione: .
También puede seleccionar uno de estos países/idiomas:
Cómo obtener el mejor rendimiento
Seleccione China (en idioma chino o inglés) para obtener el mejor rendimiento. Los sitios web de otros países no están optimizados para ser accedidos desde su ubicación geográfica.
América
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)