dddtreecfs
Extract dual-tree/double-density wavelet coefficients or projections
Syntax
Description
out = dddtreecfs(outputtype,wt,outputspec,outputindices)wt. If outputtype equals 'e', out contains
wavelet or scaling coefficients. If outputtype equals 'r', out contains
wavelet or scaling subspace projections (reconstructions).
out = dddtreecfs(outputtype,wt,outputspec,outputindices,'plot')'plot' option anywhere after
the wt input.
Examples
Obtain the complex dual-tree wavelet transform of the 1-D noisy Doppler signal. Reconstruct an approximation based on the level-three detail coefficients in multiple ways.
Load the noisy Doppler signal. Obtain the complex dual-tree transform down to level 3.
load noisdopp; wt = dddtree('cplxdt',noisdopp,3,'dtf1')
wt = struct with fields:
type: 'cplxdt'
level: 3
filters: [1×1 struct]
cfs: {[1×512×2 double] [1×256×2 double] [1×128×2 double] [1×128×2 double]}
Plot a reconstruction of the original signal based on the level-three detail coefficients with outputspec set to 'scale'.
xr = dddtreecfs('r',wt,'scale',{3},'plot');
![Figure CPLXDT contains an axes object. The axes object with title Nodes of Tree: [3], xlabel Reconstructed signal contains an object of type line.](../../examples/wavelet/win64/Reconstructionfrom1DComplexDualTreeWaveletTransformExample_01.png)
The output xr is a 1-by-1 cell array. Generate the same reconstruction by using 'cumind' and the level-three tree nodes. The first element of each vector in the cell array denotes the level, and the second element denotes the tree. Confirm the reconstructions are identical.
outputindices = {[3 1];[3 2]};
xr2 = dddtreecfs('r',wt,'cumind',outputindices);
max(abs(xr2-xr{1}))ans = 0
The output xr2 is the same datatype as the original signal.
Load the noisy Doppler signal. Obtain the complex dual-tree transform down to level 3.
load noisdopp; wt = dddtree('cplxdt',noisdopp,3,'dtf1')
wt = struct with fields:
type: 'cplxdt'
level: 3
filters: [1×1 struct]
cfs: {[1×512×2 double] [1×256×2 double] [1×128×2 double] [1×128×2 double]}
Create a cell array of vectors to obtain the second- and third-level detail coefficients from each of the wavelet filter bank trees.
outputindices = {[2 1]; [2 2]; [3 1]; [3 2]};The first element of each vector in the cell array denotes the level, or stage. The second element denotes the tree.
Extract the detail coefficients.
detailCoeffs = dddtreecfs('e',wt,'ind',outputindices,'plot');

The output detailCoeffs is a 1-by-4 cell array. The cell array elements contain the wavelet coefficients corresponding to the elements in outputindices. For example, confirm detailCoeffs{1} contains the level-two detail coefficients from the first tree.
max(abs(wt.cfs{2}(1,:,1)-detailCoeffs{1}))ans = 0
Load the noisy Doppler signal. Obtain the complex dual-tree transform down to level 3.
load noisdopp; wt = dddtree('cplxdt',noisdopp,3,'dtf1');
Create a cell array of vectors to obtain the second- and third-level detail coefficients from each of the wavelet filter bank trees.
outputindices = {[2 1]; [2 2]; [3 1];[3 2]};The first element of each vector in the cell array denotes the level, or stage. The second element denotes the tree.
Create a structure array identical to the wt output of dddtree with all the coefficients equal to zero except the second- and third-level detail coefficients.
out = dddtreecfs('e',wt,'cumind',outputindices,'plot');

Generate a reconstruction based on the second- and third-level detail coefficients.
xr = idddtree(out);
Generate two reconstructions, based on the second- and third-level detail coefficients. Confirm the sum of the two reconstructions is identical to xr.
xr2 = dddtreecfs('r',wt,'scale',{2;3}); max(abs(xr-(xr2{1}+xr2{2})))
ans = 8.8818e-16
Use the complex dual-tree wavelet transform to isolate diagonal features in an image at +45 and –45 degrees.
Load and display the xbox image.
load xbox
imagesc(xbox)
Obtain the complex dual-tree wavelet transform down to level 3.
fdf = dtfilters('FSfarras'); df = dtfilters('qshift10'); wt = dddtree2('cplxdt',xbox,3,fdf,df);
Isolate the +45 and -45 diagonal image features in the level-one wavelet coefficients. Do this by creating a cell array of vectors specifying the tree nodes containing the diagonal details. The first element in the vector specifies the level. The three remaining elements specify the orientation, wavelet tree, and real and imaginary parts, respectively (see dddtree2).
outputindices = {[1 3 1 1];[1 3 1 2];[1 3 2 1];[1 3 2 2]};
out = dddtreecfs('e',wt,'ind',outputindices,'plot');
This example shows how the analysis coefficients are distributed, depending on the transform, in the tree output of dddtree and dddtree2.
1-D Wavelet Transforms
Load in the noisy Doppler signal. Generate a four-level wavelet decomposition of the signal for each type of transform. Depending on the transform, different dimensions of the coefficient arrays correspond to orientation, wavelet tree, or real and imaginary parts.
Critically Sampled Discrete Wavelet Transform
load noisdopp wt = dddtree('dwt',noisdopp,4,'sym4')
wt = struct with fields:
type: 'dwt'
level: 4
filters: [1×1 struct]
cfs: {[1×512 double] [1×256 double] [1×128 double] [1×64 double] [1×64 double]}
This is the usual nonredundant discrete wavelet transform. The first four elements of wt.cfs are the wavelet coefficients. The fifth element are the scaling coefficients.
Double-Density Wavelet Transform
wt = dddtree('ddt',noisdopp,4,'filters1')
wt = struct with fields:
type: 'ddt'
level: 4
filters: [1×1 struct]
cfs: {[1×512×2 double] [1×256×2 double] [1×128×2 double] [1×64×2 double] [1×64 double]}
The third dimension of the 3-D wavelet coefficient arrays corresponds to the tree. The fifth element are the scaling coefficients.
Dual-Tree Complex Wavelet Transform
wt = dddtree('cplxdt',noisdopp,4,'dtf1')
wt = struct with fields:
type: 'cplxdt'
level: 4
filters: [1×1 struct]
cfs: {[1×512×2 double] [1×256×2 double] [1×128×2 double] [1×64×2 double] [1×64×2 double]}
The third dimension of all the 3-D arrays in cfs corresponds to the real and imaginary parts. The first four elements of cfs are the wavelet coefficients, and cfs{5} are the scaling coefficients.
Reconstruct signals from the coefficients at the tree nodes [1 1], [5 2], [3 1], and [4 2]. Plot the signals. The output is a cell array containing the reconstructions. The reconstructions are the same length as the original signal.
outputindices = {[1 1];[5 2];[3 1];[4 2]};
XR = dddtreecfs('r',wt,'plot','ind',outputindices);
Extract and plot the coefficients used to reconstruct the signals. The output is a cell array containing the coefficients of respective length: 512, 64, 128, and 64.
XR = dddtreecfs('e',wt,'plot','ind',outputindices);

Now use 'cumind' instead of 'ind'. The output XR is a signal of length 1024 in the first case, and a 'cplxdt' dual-tree in the second one.
XR = dddtreecfs('r',wt,'plot','cumind',outputindices);
![Figure CPLXDT contains an axes object. The axes object with title Nodes of Tree: [1 1] , [5 2] , [3 1] , [4 2], xlabel Reconstructed signal contains an object of type line.](../../examples/wavelet/win64/DistributionOfAnalysisCoefficientsInWaveletTreeStructureExample_03.png)
XR = dddtreecfs('e',wt,'plot','cumind',outputindices);

Double-Density Dual-Tree Complex Wavelet Transform
wt = dddtree('cplxdddt',noisdopp,4,'dddtf1')
wt = struct with fields:
type: 'cplxdddt'
level: 4
filters: [1×1 struct]
cfs: {[1×512×2×2 double] [1×256×2×2 double] [1×128×2×2 double] [1×64×2×2 double] [1×64×2 double]}
The third dimension of the 4-D wavelet coefficient arrays corresponds to the tree. The fourth dimension in the 4-D wavelet coefficient arrays and third dimension in the 3-D scaling coefficients array corresponds to the real and imaginary parts.
2-D Wavelet Transforms
Load in the 256-by-256 mask image. Generate a two-level wavelet decomposition of the image for each type of transform. Observe the dimensions of the output coefficients.
Critically Sampled Discrete Wavelet Transform
load mask im = X; wt = dddtree2('dwt',im,3,'sym4')
wt = struct with fields:
type: 'dwt'
level: 3
filters: [1×1 struct]
cfs: {[128×128×3 double] [64×64×3 double] [32×32×3 double] [32×32 double]}
sizes: [10×2 double]
This is the usual nonredundant 2-D discrete wavelet transform. The third dimension in the 3-D wavelet coefficient arrays corresponds to the orientation. The scaling coefficients are the last element of cfs.
Real Oriented Dual-Tree Wavelet Transform
wt = dddtree2('realdt',im,3,'dtf1')
wt = struct with fields:
type: 'realdt'
level: 3
filters: [1×1 struct]
cfs: {[128×128×3×2 double] [64×64×3×2 double] [32×32×3×2 double] [32×32×2 double]}
sizes: [11×2 double]
The fourth dimension in the 4-D wavelet coefficient arrays and third dimension in the 3-D scaling coefficients array correspond to the tree. The third dimension in the 4-D wavelet coefficient arrays corresponds to orientation.
Complex Oriented Dual-Tree Wavelet Transform
wt = dddtree2('cplxdt',im,3,'dtf1')
wt = struct with fields:
type: 'cplxdt'
level: 3
filters: [1×1 struct]
cfs: {[5-D double] [5-D double] [5-D double] [32×32×2×2 double]}
sizes: [11×2 double]
[size(wt.cfs{1});size(wt.cfs{2});size(wt.cfs{3})]ans = 3×5
128 128 3 2 2
64 64 3 2 2
32 32 3 2 2
The third dimension of the 5-D wavelet coefficient arrays represents the orientation. The fourth dimension in the 5-D arrays and third dimension in the 4-D scaling coefficients array represents the tree. The fifth dimension in the 5-D arrays and fourth dimension in the 4-D array represents the real and imaginary parts.
Double-Density Wavelet Transform
wt = dddtree2('ddt',im,3,'filters1')
wt = struct with fields:
type: 'ddt'
level: 3
filters: [1×1 struct]
cfs: {[128×128×8 double] [64×64×8 double] [32×32×8 double] [32×32 double]}
sizes: [26×2 double]
The third dimension in the 3-D wavelet coefficient arrays represents the orientation.
Real Oriented Double-Density Wavelet Transform
wt = dddtree2('realdddt',im,3,'self1')
wt = struct with fields:
type: 'realdddt'
level: 3
filters: [1×1 struct]
cfs: {[128×128×8×2 double] [64×64×8×2 double] [32×32×8×2 double] [32×32×2 double]}
sizes: [26×2 double]
The third dimension in the 4-D wavelet coefficient arrays represents the orientation. The fourth dimension in the 4-D arrays and third dimension in the 3-D scaling coefficients array represent the tree.
Complex Oriented Double-Density Wavelet Transform
wt = dddtree2('cplxdddt',im,3,'self1')
wt = struct with fields:
type: 'cplxdddt'
level: 3
filters: [1×1 struct]
cfs: {[5-D double] [5-D double] [5-D double] [32×32×2×2 double]}
sizes: [26×2 double]
[size(wt.cfs{1}) ; size(wt.cfs{2}) ; size(wt.cfs{3})]ans = 3×5
128 128 8 2 2
64 64 8 2 2
32 32 8 2 2
The third dimension of the 5-D wavelet coefficient arrays represents the orientation. The fourth dimension in the 5-D arrays and third dimension in the 4-D scaling coefficients array represents the tree. The fifth dimension in the 5-D arrays and fourth dimension in the 4-D array represents the real and imaginary parts.
Reconstruct and plot two images based on the second-level detail coefficients and scaling coefficients, respectively.
XR = dddtreecfs('r',wt,'plot','scale',{2;4});
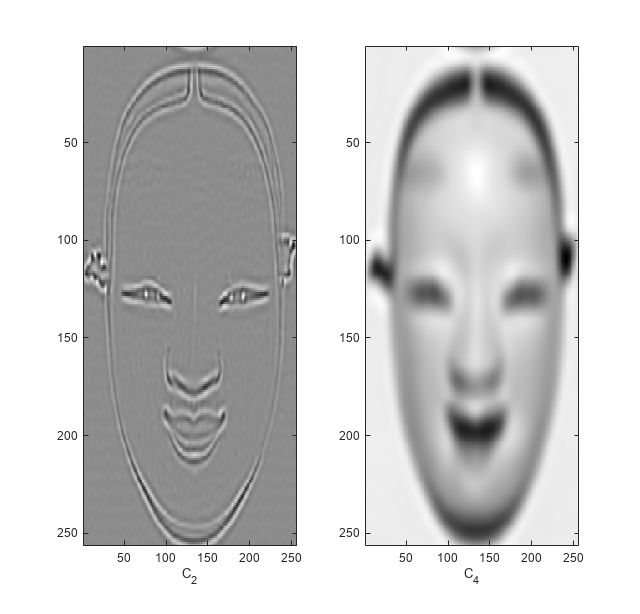
The output XR is a cell array containing both 256-by-256 images.
Extract the coefficients used to produce the two images. The output is a cell array containing two dual-tree structures, one for each specified scale.
XR = dddtreecfs('e',wt,'scale',{2;4}); XR{1}
ans = struct with fields:
type: 'cplxdddt'
level: 3
filters: [1×1 struct]
cfs: {[5-D double] [5-D double] [5-D double] [32×32×2×2 double]}
sizes: [26×2 double]
XR{2}ans = struct with fields:
type: 'cplxdddt'
level: 3
filters: [1×1 struct]
cfs: {[5-D double] [5-D double] [5-D double] [32×32×2×2 double]}
sizes: [26×2 double]
Confirm the only nonzero coefficients in each structure contained in XR are the level-two wavelet coefficients and scaling coefficients, respectively.
dtInd = 1;
[max(abs(XR{dtInd}.cfs{1}(:)));max(abs(XR{dtInd}.cfs{2}(:)));...
max(abs(XR{dtInd}.cfs{3}(:)));max(abs(XR{dtInd}.cfs{4}(:)))]ans = 4×1
0
143.9924
0
0
dtInd = 2;
[max(abs(XR{dtInd}.cfs{1}(:)));max(abs(XR{dtInd}.cfs{2}(:)));...
max(abs(XR{dtInd}.cfs{3}(:)));max(abs(XR{dtInd}.cfs{4}(:)))]ans = 4×1
103 ×
0
0
0
1.0545
Use 'ind' to reconstruct and display the four images based on the four lowpass components, respectively.
outputindices = {[4 1 1];[4 2 1];[4 1 2];[4 2 2]};
XR = dddtreecfs('r',wt,'plot','ind',outputindices);
The output XR is a cell array containing the four images. Each image is 256-by-256. Display the coefficients used to reconstruct the images.
XR = dddtreecfs('e',wt,'plot','ind',outputindices);

The output XR is a cell array containing the four lowpass components. Each component is 32-by-32.
Input Arguments
Output type, specified as 'e' or 'r'.
Use 'e' to obtain the scaling or wavelet coefficients.
Use 'r' to obtain a projection, or reconstruction,
onto the appropriate scaling or wavelet subspace.
Wavelet transform, specified as a structure. The structure array
is the output of dddtree or dddtree2.
Output specification, specified as one of 'lowpass', 'scale', 'ind',
or 'cumind'. The output specifications are defined
as follows:
'lowpass'— Outputs the lowpass, or scaling, coefficients or a signal/image approximation based on the scaling coefficients. If you set the output specification to'lowpass', do not specifyoutputindices. If theoutputtypeis'e',outis a structure array with fields identical to the input structure arraywtexcept that all wavelet (detail) coefficients are equal to zero. If theoutputtypeis'r',outis a signal or image approximation based on the scaling coefficients. The signal or image approximation is equal in size to the original input todddtreeordddtree2.'scale'— Outputs the coefficients or a signal/image approximation based on the scales specified inoutputindices. If theoutputtypeis'e',outis a cell array of structure arrays. The fields of the structure arrays inoutare identical to the fields of the input structure arraywt. The coefficients in thecfsfield are all equal to zero except the coefficients corresponding to the scales inoutputindices. If theoutputtypeis'r',outis a signal or image approximation based on the scales inoutputindices. The signal or image approximation is equal in size to the original input todddtreeordddtree2.'ind'— Outputs the coefficients or a signal/image approximation based on the tree-position indices specified inoutputindices. If theoutputtypeis'e',outis a cell array of vectors or matrices containing the coefficients specified by the tree-position indices inoutputindices. If theoutputtypeis'r',outis a cell array of vectors or matrices containing signal or image approximations based on the corresponding tree-position indices inoutputindices.'cumind'— Outputs the coefficients or a signal/image approximation based on the tree-position indices specified inoutputindices. If theoutputtypeis'e',outis a structure array. The fields of the structure array are identical to the fields of the input structure arraywt. The coefficients in thecfsfield are all equal to zero except the coefficients corresponding to the tree positions inoutputindices. If theoutputtypeis'r',outis a signal or image approximation based on the coefficients corresponding to the tree-position indices inoutputindices.
Example: 'ind',{[1 1]; [1 2]}
Output indices, specified as a cell array with scalar or vector elements. If
outputspec equals 'scale', a
scalar element selects the corresponding element in the
cfs field of wt. If
outputspec equals 'ind' or
'cumind', the elements of
outputspec are row vectors. The first element of
the row vector corresponds to the element in the cfs
field of wt. Subsequent elements in the row vector
correspond to the indices of the array contained in the cell array element.
For a description of the subsequent elements, see Distribution of Analysis Coefficients in Wavelet Tree Structure. For
more information, see dddtree and dddtree2.
Example: 'scale',{1;2;3}
Output Arguments
Signal or image reconstruction or coefficients, returned as a vector,
matrix, structure array, cell array of vectors or matrices, or cell array of
structure arrays. The form of out depends on the value
of outputspec and
outputindices.
Version History
Introduced in R2013b
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleccione un país/idioma
Seleccione un país/idioma para obtener contenido traducido, si está disponible, y ver eventos y ofertas de productos y servicios locales. Según su ubicación geográfica, recomendamos que seleccione: .
También puede seleccionar uno de estos países/idiomas:
Cómo obtener el mejor rendimiento
Seleccione China (en idioma chino o inglés) para obtener el mejor rendimiento. Los sitios web de otros países no están optimizados para ser accedidos desde su ubicación geográfica.
América
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)