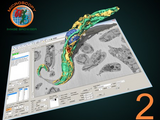
MIB2 is an update package for segmentation of multi-dimensional (2D-4D) microscopy datasets
Ahora está siguiendo esta publicación
- Verá actualizaciones en las notificaciones de contenido en seguimiento.
- Podrá recibir correos electrónicos, en función de las preferencias de comunicación que haya establecido.
- Support 2D-4D datasets (x,y,c,z,t)
- Up to 9 simultaneously opened datasets
- Bounding box for each dataset
- Extendible via plugins
- Log of performed actions
- Customizable undo system
- Customizable keyboard shortcuts
- Colorblind friendly color schemes
- Regions of interests
- Virtual stacking mode for working with datasets that are larger than available memory
- Batch processing mode
- Direct import/export with Matlab , Fiji , Imaris and system clipboard
- Direct import from Omero server and URL links
- Load and save to TIF, Amira Mesh, JPG, Fiji BigDataViewer, HDF5, MRC, NRRD, PNG formats
- Load up to 100 different image and video formats
- Microsoft Excel (export) for quantification
- Rename and shuffle tool for unbiased classification and segmentation
- Objects: Area (2D/3D)
- Objects: ConvexArea (2D)
- Objects: Curve Length (2D, pixels and image units)
- Objects: Eccentricity (2D)
- Objects: Equatorial Eccentricity (3D)
- Objects: Equiv Diameter (2D)
- Objects: Euler number (2D)
- Objects: Extent (2D)
- Objects: Filled area (2D/3D)
- Objects: Holes area (2D/3D)
- Objects: Length between end points (2D/3D)
- Objects: Major axis length (2D/3D)
- Objects: Meridional Eccentricity (3D)
- Objects: Orientation (2D)
- Objects: Perimeter (2D)
- Objects: Second axis length (2D/3D)
- Objects: Solidity (2D)
- Objects: Third axis length (3D)
- Intensity: Correlation (2D/3D)
- Intensity: Maximal (2D/3D)
- Intensity: Mean (2D/3D)
- Intensity: Minimal (2D/3D)
- Intensity: Standard deviation (2D/3D)
- Intensity: Sum (2D/3D)
- Angles
- Caliper
- Circle, radius
- Freehand distance and intensity profile
- Linear distance and intensity profile
- Polyline distance and intensity profile
- Stereology
- Wound healing assay
- 3D ball (3D)
- 3D lines (3D)
- Annotations with values
- Brush tool (2D)
- Brush tool for 2D superpixels (SLIC , Watershed )
- Black and White Thresholding tool (global, local, adaptive; 2D/3D)
- Deep convolutional neural networks training for segmentation (2D, 2.5D, 3D, 2D patch-wise)
- Dilate (2D/3D, difference)
- Drag & Drop
- Erode (2D/3D, difference)
- Fill holes (2D/3D)
- Frame selection tool
- Frangi tubular filter (2D/3D)
- Graphcut based semi-automatic segmentation(2D/3D) ,
- Lasso tool (2D/3D)
- Magic Wand tool (2D/3D)
- Membrane Click Tracker tool (2D/3D)
- Morphological operations (branch points, diagonal fill, end points, skeleton, spur, thin, ultimate erosion)
- Object Picker (2D/3D)
- Quantification Filtering (2D/3D)
- Random Forest Classifier (2D/3D)
- Shape and Line Interpolation (3D)
- Smooth (2D/3D)
- Segment-anything model v1 and v2 for interactive 2D and 3D segmentation
- Spot tool (2D/3D)
- Watershed for automatic image segmentation and object separation (2D/3D)
- Add frame around the dataset
- Alignment
- Brightness, Contrast, Gamma adjustments
- Chop and re-chop large dataset to smaller volumes
- Content-aware fill
- Contrast-limited adaptive histogram equalization
- Color mode change (depth, color type)
- Color channel operations (add, copy, delete, invert, rotate, shift, swap)
- Crop , Resize , Flip , Rotate , Transpose
- Crop 2D/3D objects to files
- Debris removal
- Image arithmetics
- Image filters
- Intensity normalization in Z/T (complete slice, masked areas, background shift)
- Intensity replacement within selected areas
- Invert
- Manipulations with slices: insert, copy, delete
- Intensity projections and focus stacking
- Morphological operations
- Orthoslices (XY, ZX, ZY planes)
- Volume Rendering (hardware)
- Volume Rendering (software)
- Models with Matlab isosurfaces
- Models and volumes with Fiji 3D viewer
- Models and volumes with Imaris
- Export models to IMOD
- Export models to Amira
- Export models to 3D Slicer
- Export models and volumes to Matlab Volume Viewer
- Export models in STL format
Citar como
Ilya Belevich (2026). Microscopy Image Browser 2 (MIB2) (https://github.com/Ajaxels/MIB2), GitHub. Recuperado .
Belevich, Ilya, et al. “Microscopy Image Browser: A Platform for Segmentation and Analysis of Multidimensional Datasets.” PLOS Biology, vol. 14, no. 1, Public Library of Science (PLoS), Jan. 2016, p. e1002340, doi:10.1371/journal.pbio.1002340.
Belevich, Ilya, and Eija Jokitalo. DeepMIB: User-Friendly and Open-Source Software for Training of Deep Learning Network for Biological Image Segmentation. Cold Spring Harbor Laboratory, July 2020, doi:10.1101/2020.07.13.200105.
Agradecimientos
Inspirado por: view3d.m, IMAGEVIEWER, findjobj - find java handles of Matlab graphic objects, 3D Euclidean Distance Transform for Variable Data Aspect Ratio, Region Adjacency Graph (RAG), stlwrite - write ASCII or Binary STL files, maxflow, Viewer3D, export_fig, Hessian based Frangi Vesselness filter, Accurate Fast Marching, Fast/Robust Template Matching, imgaussian, Highly portable JSON-input parser, Jann5s/measuretool, Render RGB text over RGB or Grayscale Image, improved xlswrite.m, xml2struct, struct2xml, Hardware accelerated 3D viewer for MATLAB, mdrohmann/mtocpp, xlwrite: Generate XLS(X) files without Excel on Mac/Linux/Win, drifty_shifty_deluxe.m, regionprops3
Información general
- Versión 2.9100 (91,5 MB)
-
Ver licencia en GitHub
Compatibilidad con la versión de MATLAB
- Compatible con cualquier versión desde R2014b
Compatibilidad con las plataformas
- Windows
- macOS
- Linux
No se pueden descargar versiones que utilicen la rama predeterminada de GitHub
| Versión | Publicado | Notas de la versión | Action |
|---|---|---|---|
| 2.9100 | tons of changes: https://mib.helsinki.fi/downloads.html |
||
| 2.8020 | bug fixes:
|
||
| 2.8002 | bug fix |
||
| 2.8001 | updated description |
||
| 2.80 | The update includes multiple improvements especially in training and segmentation of 2D convolutional neural networks with DeepMIB.
|
||
| 2.70 | - deep learning with DeepMIB to train deep convolutional networks for image segmentation
|
||
| 2.60 | Batch processing mode demo
|
||
| 2.1.0.0 | added a demo image |