margin
Classification margins for Gaussian kernel classification model
Description
m = margin(Mdl,Tbl,ResponseVarName)Mdl using the predictor data in table
Tbl and the class labels in
Tbl.ResponseVarName.
Examples
Load the ionosphere data set. This data set has 34 predictors and 351 binary responses for radar returns, either bad ('b') or good ('g').
load ionospherePartition the data set into training and test sets. Specify a 30% holdout sample for the test set.
rng('default') % For reproducibility Partition = cvpartition(Y,'Holdout',0.30); trainingInds = training(Partition); % Indices for the training set testInds = test(Partition); % Indices for the test set
Train a binary kernel classification model using the training set.
Mdl = fitckernel(X(trainingInds,:),Y(trainingInds));
Estimate the training-set margins and test-set margins.
mTrain = margin(Mdl,X(trainingInds,:),Y(trainingInds)); mTest = margin(Mdl,X(testInds,:),Y(testInds));
Plot both sets of margins using box plots.
boxplot([mTrain; mTest],[zeros(size(mTrain,1),1); ones(size(mTest,1),1)], ... 'Labels',{'Training set','Test set'}); title('Training-Set and Test-Set Margins')
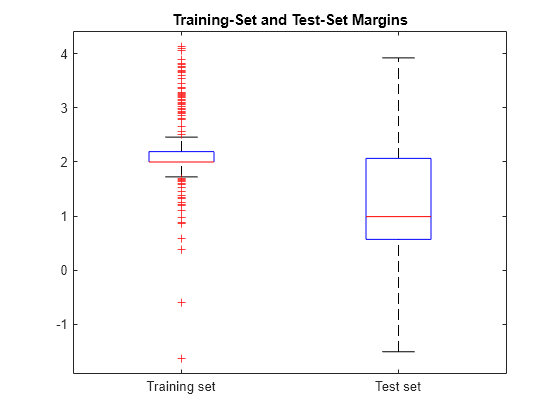
The margin distribution of the training set is situated higher than the margin distribution of the test set.
Perform feature selection by comparing test-set margins from multiple models. Based solely on this criterion, the classifier with the larger margins is the better classifier.
Load the ionosphere data set. This data set has 34 predictors and 351 binary responses for radar returns, either bad ('b') or good ('g').
load ionospherePartition the data set into training and test sets. Specify a 15% holdout sample for the test set.
rng('default') % For reproducibility Partition = cvpartition(Y,'Holdout',0.15); trainingInds = training(Partition); % Indices for the training set XTrain = X(trainingInds,:); YTrain = Y(trainingInds); testInds = test(Partition); % Indices for the test set XTest = X(testInds,:); YTest = Y(testInds);
Randomly choose 10% of the predictor variables.
p = size(X,2); % Number of predictors
idxPart = randsample(p,ceil(0.1*p));Train two binary kernel classification models: one that uses all of the predictors, and one that uses the random 10%.
Mdl = fitckernel(XTrain,YTrain); PMdl = fitckernel(XTrain(:,idxPart),YTrain);
Mdl and PMdl are ClassificationKernel models.
Estimate the test-set margins for each classifier.
fullMargins = margin(Mdl,XTest,YTest); partMargins = margin(PMdl,XTest(:,idxPart),YTest);
Plot the distribution of the margin sets using box plots.
boxplot([fullMargins partMargins], ... 'Labels',{'All Predictors','10% of the Predictors'}); title('Test-Set Margins')
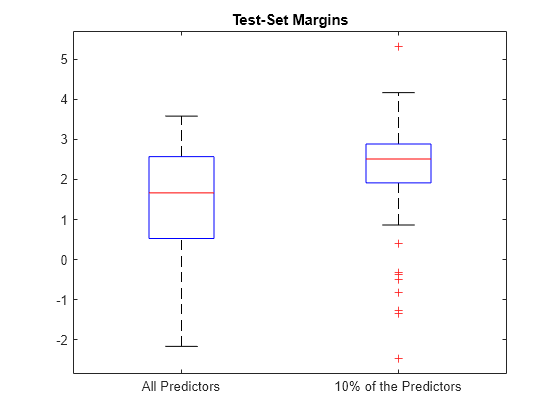
The margin distribution of PMdl is situated higher than the margin distribution of Mdl. Therefore, the PMdl model is the better classifier.
Input Arguments
Binary kernel classification model, specified as a ClassificationKernel model object. You can create a
ClassificationKernel model object using fitckernel.
Class labels, specified as a categorical, character, or string array; logical or numeric vector; or cell array of character vectors.
The data type of
Ymust be the same as the data type ofMdl.ClassNames. (The software treats string arrays as cell arrays of character vectors.)The distinct classes in
Ymust be a subset ofMdl.ClassNames.If
Yis a character array, then each element must correspond to one row of the array.The length of
Ymust be equal to the number of observations inXorTbl.
Data Types: categorical | char | string | logical | single | double | cell
Sample data used to train the model, specified as a table. Each row of
Tbl corresponds to one observation, and each column corresponds
to one predictor variable. Optionally, Tbl can contain additional
columns for the response variable and observation weights. Tbl must
contain all the predictors used to train Mdl. Multicolumn variables
and cell arrays other than cell arrays of character vectors are not allowed.
If Tbl contains the response variable used to train Mdl, then you do not need to specify ResponseVarName or Y.
If you train Mdl using sample data contained in a table, then the input
data for margin must also be in a table.
Response variable name, specified as the name of a variable in Tbl. If Tbl contains the response variable used to train Mdl, then you do not need to specify ResponseVarName.
If you specify ResponseVarName, then you must specify it as a character
vector or string scalar. For example, if the response variable is stored as
Tbl.Y, then specify ResponseVarName as
'Y'. Otherwise, the software treats all columns of
Tbl, including Tbl.Y, as predictors.
The response variable must be a categorical, character, or string array; a logical or numeric vector; or a cell array of character vectors. If the response variable is a character array, then each element must correspond to one row of the array.
Data Types: char | string
Output Arguments
Classification margins, returned as an n-by-1
numeric column vector, where n is the number of
observations in X.
More About
The classification margin for binary classification is, for each observation, the difference between the classification score for the true class and the classification score for the false class.
The software defines the classification margin for binary classification as
x is an observation. If the true label of x is the positive class, then y is 1, and –1 otherwise. f(x) is the positive-class classification score for the observation x. The classification margin is commonly defined as m = yf(x).
If the margins are on the same scale, then they serve as a classification confidence measure. Among multiple classifiers, those that yield greater margins are better.
For kernel classification models, the raw classification score for classifying the observation x, a row vector, into the positive class is defined by
is a transformation of an observation for feature expansion.
β is the estimated column vector of coefficients.
b is the estimated scalar bias.
The raw classification score for classifying x into the negative class is −f(x). The software classifies observations into the class that yields a positive score.
If the kernel classification model consists of logistic regression learners, then the
software applies the 'logit' score transformation to the raw
classification scores (see ScoreTransform).
Extended Capabilities
The
margin function supports tall arrays with the following usage
notes and limitations:
margindoes not support talltabledata.
For more information, see Tall Arrays.
This function fully supports GPU arrays. For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2017bmargin fully supports GPU arrays.
See Also
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleccione un país/idioma
Seleccione un país/idioma para obtener contenido traducido, si está disponible, y ver eventos y ofertas de productos y servicios locales. Según su ubicación geográfica, recomendamos que seleccione: .
También puede seleccionar uno de estos países/idiomas:
Cómo obtener el mejor rendimiento
Seleccione China (en idioma chino o inglés) para obtener el mejor rendimiento. Los sitios web de otros países no están optimizados para ser accedidos desde su ubicación geográfica.
América
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)