GapEvaluation
Gap criterion clustering evaluation object
Description
GapEvaluation is an object consisting of sample data (X), clustering data (OptimalY), and gap criterion values
(CriterionValues) used to
evaluate the optimal number of clusters (OptimalK). The gap criterion values
correspond to the difference ExpectedLogW –
LogW, where W is the within-cluster dispersion,
ExpectedLogW is determined by Monte Carlo sampling from a reference
distribution, and LogW is computed from the sample data. The optimal
number of clusters corresponds to the solution with the largest local or global gap value
within a tolerance range (SearchMethod). For
more information, see Gap Value.
Creation
Create a gap criterion clustering evaluation object by using the evalclusters function and specifying the criterion as
"gap".
You can then use compact to create a compact version of the gap
criterion clustering evaluation object. The function removes the contents of the properties
X, OptimalY, and
Missing.
Properties
Clustering Evaluation Properties
This property is read-only.
Clustering algorithm used to cluster the sample data, returned as
'kmeans', 'linkage',
'gmdistribution', or a function handle.
| Value | Description |
|---|---|
'kmeans' | Cluster the data in X using the kmeans clustering algorithm, with
EmptyAction set to "singleton" and
Replicates set to 5. |
'linkage' | Cluster the data in X using the clusterdata agglomerative
clustering algorithm, with Linkage set to
"ward". |
'gmdistribution' | Cluster the data in X using the gmdistribution Gaussian mixture
distribution algorithm, with SharedCov set to
true and Replicates set to
5. |
Data Types: char | function_handle
This property is read-only.
Name of the criterion used for clustering evaluation, returned as
'Gap'.
This property is read-only.
Criterion values, returned as a numeric vector. Each value corresponds to a proposed
number of clusters in InspectedK.
Data Types: double
This property is read-only.
Distance metric used for clustering data and computing the criterion values, returned as one of the values in this table or a function handle.
| Value | Description |
|---|---|
'sqEuclidean' | Squared Euclidean distance |
'Euclidean' | Euclidean distance |
'cityblock' | Sum of absolute differences |
'cosine' | One minus the cosine of the included angle between points (treated as vectors) |
'correlation' | One minus the sample correlation between points (treated as sequences of values) |
Data Types: char | function_handle
This property is read-only.
List of the number of proposed clusters for which to compute criterion values, returned as a positive integer vector.
Data Types: double
This property is read-only.
Optimal number of clusters, returned as a positive integer scalar.
Data Types: double
This property is read-only.
Optimal clustering solution corresponding to OptimalK, returned
as a positive integer column vector. Each row of OptimalY
represents the cluster index of the corresponding observation (or row) in
X. If you specify the clustering solutions as an input argument
to evalclusters when you create the clustering evaluation object,
or if the clustering evaluation object is compact (see compact), then OptimalY is empty.
Data Types: double
This property is read-only.
Method for selecting the optimal number of clusters, returned as
'globalMaxSE' or 'firstMaxSE'.
| Value | Description |
|---|---|
'globalMaxSE' | Evaluate each proposed number of clusters in
where K is the number of clusters, Gap(K) is the gap value for the clustering solution with K clusters, GAPMAX is the largest gap value, and SE(GAPMAX) is the standard error corresponding to the largest gap value. |
'firstMaxSE' | Evaluate each proposed number of clusters in
where K is the number of clusters, Gap(K) is the gap value for the clustering solution with K clusters, and SE(K + 1) is the standard error of the clustering solution with K + 1 clusters. |
Sample Data Properties
This property is read-only.
Natural logarithm of the within-cluster dispersion W based on
the sample data X, returned as a numeric vector.
W is the within-cluster dispersion computed using the distance
metric Distance. Each element of LogW
corresponds to a specific number of proposed clusters (an element of
InspectedK).
Data Types: double
This property is read-only.
Excluded data, returned as a logical column vector. If an element of
Missing is true, then the corresponding
observation (or row) in the data matrix X is not used in the
clustering solutions. If the clustering evaluation object is compact (see compact), then Missing is empty.
Data Types: double | logical
This property is read-only.
Number of observations in the data matrix X, ignoring
observations with missing (NaN) values, returned as a positive
integer scalar.
Data Types: double
This property is read-only.
Data used for clustering, returned as a numeric matrix. Rows correspond to
observations, and columns correspond to variables. If the clustering evaluation object
is compact (see compact), then X is
empty.
Data Types: single | double
Reference Data Properties
This property is read-only.
Number of reference data sets generated from the reference distribution
ReferenceDistribution, returned as a positive integer
scalar.
Data Types: double
This property is read-only.
Expectation of the natural logarithm of the within-cluster dispersion
W based on the generated reference data, returned as a numeric
vector. W is the within-cluster dispersion computed using the
distance metric Distance. Each element of
ExpectedLogW corresponds to a specific number of proposed
clusters (an element of InspectedK).
Data Types: double
This property is read-only.
Reference data generation method, returned as 'PCA' or
'uniform'.
| Value | Description |
|---|---|
'PCA' | Generate reference data from a uniform distribution over a box aligned
with the principal components of the data matrix
X. |
'uniform' | Generate reference data uniformly over the range of each feature in the
data matrix X. |
This property is read-only.
Standard error of the natural logarithm of the within-cluster dispersion
W with respect to the reference data, returned as a numeric
vector. W is the within-cluster dispersion computed using the
distance metric Distance. Each element of SE
corresponds to a specific number of proposed clusters (an element of
InspectedK).
Data Types: double
This property is read-only.
Standard deviation of the natural logarithm of the within-cluster dispersion
W with respect to the reference data, returned as a numeric
vector. W is the within-cluster dispersion computed using the
distance metric Distance. Each element of
StdLogW corresponds to a specific number of proposed clusters
(an element of InspectedK).
Data Types: double
Object Functions
Examples
Evaluate the optimal number of clusters using the gap clustering evaluation criterion.
Load the fisheriris data set. The data contains length and width measurements from the sepals and petals of three species of iris flowers.
load fisheririsEvaluate the optimal number of clusters based on the gap criterion values. Cluster the data using kmeans.
rng("default") % For reproducibility evaluation = evalclusters(meas,"kmeans","gap","KList",1:6)
evaluation =
GapEvaluation with properties:
NumObservations: 150
InspectedK: [1 2 3 4 5 6]
CriterionValues: [0.0720 0.5928 0.8762 1.0114 1.0534 1.0720]
OptimalK: 5
Properties, Methods
The OptimalK value indicates that, based on the gap criterion, the optimal number of clusters is five.
Plot the gap criterion values for each number of clusters tested.
plot(evaluation)
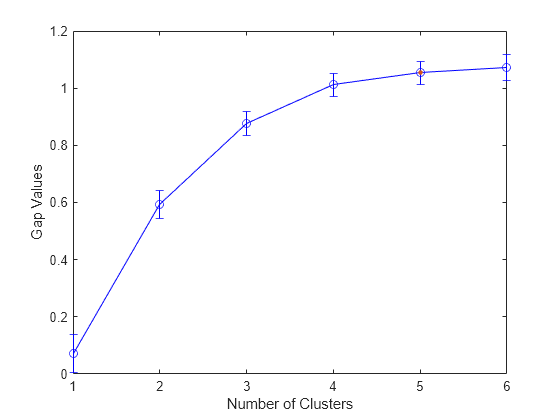
Based on the plot, the maximum value of the gap criterion occurs at six clusters. However, the value at five clusters is within one standard error of the maximum, so the suggested optimal number of clusters is five.
Create a grouped scatter plot to examine the relationship between petal length and width. Group the data by the suggested clusters.
PetalLength = meas(:,3);
PetalWidth = meas(:,4);
clusters = evaluation.OptimalY;
gscatter(PetalLength,PetalWidth,clusters,[],"xod^*");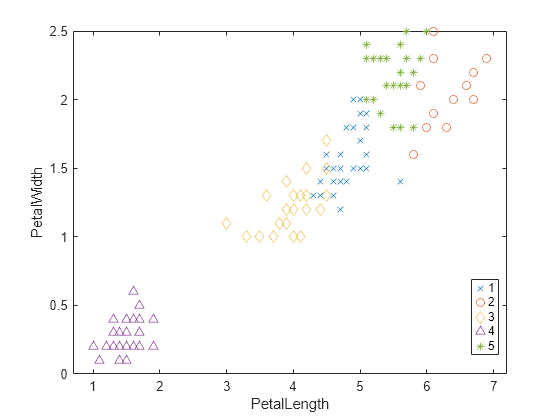
The plot shows cluster 4 in the lower-left corner, completely separated from the other four clusters. Cluster 4 contains flowers with the smallest petal widths and lengths. Cluster 2 is in the upper-right corner, and contains flowers with the largest petal widths and lengths. Cluster 5 is next to cluster 2, and contains flowers with similar petal widths but smaller petal lengths compared to the flowers in cluster 2. Clusters 1 and 3 are near the center of the plot, and contain flowers with measurements between the extremes.
More About
A common graphical approach to clustering evaluation involves plotting an error measurement versus several proposed numbers of clusters, and locating the “elbow” of this plot. The “elbow” occurs at the most dramatic decrease in error measurement. The gap criterion formalizes this approach by estimating the “elbow” location as the number of clusters with the largest gap value. Therefore, under the gap criterion, the optimal number of clusters corresponds to the solution with the largest local or global gap value within a tolerance range.
The gap value is defined as
where n is the sample size, k is the number of clusters being evaluated, and Wk is the pooled within-cluster dispersion measurement
where nr is the number of data points in cluster r, and Dr is the sum of the pairwise distances for all points in cluster r.
The expected value is determined by Monte Carlo sampling from a reference distribution, and
log(Wk) is computed from
the sample data.
The gap value is defined even for clustering solutions that contain only one cluster, and can be used with any distance metric. However, the gap criterion is more computationally expensive than other clustering evaluation criteria, because the clustering algorithm must be applied to the reference data for each proposed clustering solution.
References
[1] Tibshirani, R., G. Walther, and T. Hastie. “Estimating the number of clusters in a data set via the gap statistic.” Journal of the Royal Statistical Society: Series B. Vol. 63, Part 2, 2001, pp. 411–423.
Version History
Introduced in R2013b
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleccione un país/idioma
Seleccione un país/idioma para obtener contenido traducido, si está disponible, y ver eventos y ofertas de productos y servicios locales. Según su ubicación geográfica, recomendamos que seleccione: .
También puede seleccionar uno de estos países/idiomas:
Cómo obtener el mejor rendimiento
Seleccione China (en idioma chino o inglés) para obtener el mejor rendimiento. Los sitios web de otros países no están optimizados para ser accedidos desde su ubicación geográfica.
América
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)