loss
Class: RegressionLinear
Regression loss for linear regression models
Description
L = loss(Mdl,Tbl,ResponseVarName)Tbl and the true
responses in Tbl.ResponseVarName.
L = loss(___,Name,Value)
Input Arguments
Linear regression model, specified as a RegressionLinear model
object. You can create a RegressionLinear model
object using fitrlinear.
Predictor data, specified as an n-by-p full or sparse matrix. This orientation of X indicates that rows correspond to individual observations, and columns correspond to individual predictor variables.
Note
If you orient your predictor matrix so that observations correspond to columns and specify 'ObservationsIn','columns', then you might experience a significant reduction in computation time.
The length of Y and the number of observations
in X must be equal.
Data Types: single | double
Sample data used to train the model, specified as a table. Each row of
Tbl corresponds to one observation, and each column corresponds
to one predictor variable. Optionally, Tbl can contain additional
columns for the response variable and observation weights. Tbl must
contain all the predictors used to train Mdl. Multicolumn variables
and cell arrays other than cell arrays of character vectors are not allowed.
If Tbl contains the response variable used to train Mdl, then you do not need to specify ResponseVarName or Y.
If you train Mdl using sample data contained in a table, then the input
data for loss must also be in a table.
Response variable name, specified as the name of a variable in
Tbl. The response variable must be a numeric
vector.
If you specify ResponseVarName, then you must specify
it as a character vector or string scalar. For example, if the response
variable is stored as Tbl.Y, then specify
ResponseVarName as 'Y'.
Otherwise, the software treats all columns of Tbl,
including Tbl.Y, as predictors.
Data Types: char | string
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Loss function, specified as the comma-separated pair consisting of
'LossFun' and a built-in loss function name or
function handle.
The following table lists the available loss functions. Specify one using its corresponding value. Also, in the table,
β is a vector of p coefficients.
x is an observation from p predictor variables.
b is the scalar bias.
Value Description 'epsiloninsensitive'Epsilon-insensitive loss: 'mse'MSE: 'epsiloninsensitive'is appropriate for SVM learners only.Specify your own function using function handle notation.
Let n be the number of observations in
X. Your function must have this signaturewhere:lossvalue =lossfun(Y,Yhat,W)The output argument
lossvalueis a scalar.You choose the function name (
lossfun).Yis an n-dimensional vector of observed responses.losspasses the input argumentYin forY.Yhatis an n-dimensional vector of predicted responses, which is similar to the output ofpredict.Wis an n-by-1 numeric vector of observation weights.
Specify your function using
'LossFun',@.lossfun
Data Types: char | string | function_handle
Since R2023b
Predicted response value to use for observations with missing predictor values,
specified as "median", "mean",
"omitted", or a numeric scalar.
| Value | Description |
|---|---|
"median" | loss uses the median of the observed
response values in the training data as the predicted response value for
observations with missing predictor values. |
"mean" | loss uses the mean of the observed
response values in the training data as the predicted response value for
observations with missing predictor values. |
"omitted" | loss excludes observations with missing
predictor values from the loss computation. |
| Numeric scalar | loss uses this value as the predicted
response value for observations with missing predictor values. |
If an observation is missing an observed response value or an observation weight, then
loss does not use the observation in the loss
computation.
Example: PredictionForMissingValue="omitted"
Data Types: single | double | char | string
Predictor data observation dimension, specified as 'rows' or
'columns'.
Note
If you orient your predictor matrix so that observations correspond to columns and
specify 'ObservationsIn','columns', then you might experience a
significant reduction in computation time. You cannot specify
'ObservationsIn','columns' for predictor data in a
table.
Data Types: char | string
Observation weights, specified as the comma-separated pair consisting
of 'Weights' and a numeric vector or the name of a
variable in Tbl.
If you specify
Weightsas a numeric vector, then the size ofWeightsmust be equal to the number of observations inXorTbl.If you specify
Weightsas the name of a variable inTbl, then the name must be a character vector or string scalar. For example, if the weights are stored asTbl.W, then specifyWeightsas'W'. Otherwise, the software treats all columns ofTbl, includingTbl.W, as predictors.
If you supply weights, loss computes the weighted
regression loss and normalizes Weights to sum to
1.
Data Types: double | single
Output Arguments
Note
If Mdl.FittedLoss is 'mse',
then the loss term in the objective function is half of the MSE. loss returns
the MSE by default. Therefore, if you use loss to
check the resubstitution (training) error, then there is a discrepancy
between the MSE and optimization results that fitrlinear returns.
Examples
Simulate 10000 observations from this model
is a 10000-by-1000 sparse matrix with 10% nonzero standard normal elements.
e is random normal error with mean 0 and standard deviation 0.3.
rng(1) % For reproducibility
n = 1e4;
d = 1e3;
nz = 0.1;
X = sprandn(n,d,nz);
Y = X(:,100) + 2*X(:,200) + 0.3*randn(n,1);Train a linear regression model. Reserve 30% of the observations as a holdout sample.
CVMdl = fitrlinear(X,Y,'Holdout',0.3);
Mdl = CVMdl.Trained{1}Mdl =
RegressionLinear
ResponseName: 'Y'
ResponseTransform: 'none'
Beta: [1000×1 double]
Bias: -0.0066
Lambda: 1.4286e-04
Learner: 'svm'
Properties, Methods
CVMdl is a RegressionPartitionedLinear model. It contains the property Trained, which is a 1-by-1 cell array holding a RegressionLinear model that the software trained using the training set.
Extract the training and test data from the partition definition.
trainIdx = training(CVMdl.Partition); testIdx = test(CVMdl.Partition);
Estimate the training- and test-sample MSE.
mseTrain = loss(Mdl,X(trainIdx,:),Y(trainIdx))
mseTrain = 0.1496
mseTest = loss(Mdl,X(testIdx,:),Y(testIdx))
mseTest = 0.1798
Because there is one regularization strength in Mdl, mseTrain and mseTest are numeric scalars.
Simulate 10000 observations from this model
is a 10000-by-1000 sparse matrix with 10% nonzero standard normal elements.
e is random normal error with mean 0 and standard deviation 0.3.
rng(1) % For reproducibility n = 1e4; d = 1e3; nz = 0.1; X = sprandn(n,d,nz); Y = X(:,100) + 2*X(:,200) + 0.3*randn(n,1); X = X'; % Put observations in columns for faster training
Train a linear regression model. Reserve 30% of the observations as a holdout sample.
CVMdl = fitrlinear(X,Y,'Holdout',0.3,'ObservationsIn','columns'); Mdl = CVMdl.Trained{1}
Mdl =
RegressionLinear
ResponseName: 'Y'
ResponseTransform: 'none'
Beta: [1000×1 double]
Bias: -0.0066
Lambda: 1.4286e-04
Learner: 'svm'
Properties, Methods
CVMdl is a RegressionPartitionedLinear model. It contains the property Trained, which is a 1-by-1 cell array holding a RegressionLinear model that the software trained using the training set.
Extract the training and test data from the partition definition.
trainIdx = training(CVMdl.Partition); testIdx = test(CVMdl.Partition);
Create an anonymous function that measures Huber loss ( = 1), that is,
where
is the residual for observation j. Custom loss functions must be written in a particular form. For rules on writing a custom loss function, see the 'LossFun' name-value pair argument.
huberloss = @(Y,Yhat,W)sum(W.*((0.5*(abs(Y-Yhat)<=1).*(Y-Yhat).^2) + ...
((abs(Y-Yhat)>1).*abs(Y-Yhat)-0.5)))/sum(W);Estimate the training set and test set regression loss using the Huber loss function.
eTrain = loss(Mdl,X(:,trainIdx),Y(trainIdx),'LossFun',huberloss,... 'ObservationsIn','columns')
eTrain = -0.4186
eTest = loss(Mdl,X(:,testIdx),Y(testIdx),'LossFun',huberloss,... 'ObservationsIn','columns')
eTest = -0.4010
Simulate 10000 observations from this model
is a 10000-by-1000 sparse matrix with 10% nonzero standard normal elements.
e is random normal error with mean 0 and standard deviation 0.3.
rng(1) % For reproducibility
n = 1e4;
d = 1e3;
nz = 0.1;
X = sprandn(n,d,nz);
Y = X(:,100) + 2*X(:,200) + 0.3*randn(n,1);Create a set of 15 logarithmically-spaced regularization strengths from through .
Lambda = logspace(-4,-1,15);
Hold out 30% of the data for testing. Identify the test-sample indices.
cvp = cvpartition(numel(Y),'Holdout',0.30);
idxTest = test(cvp);Train a linear regression model using lasso penalties with the strengths in Lambda. Specify the regularization strengths, optimizing the objective function using SpaRSA, and the data partition. To increase execution speed, transpose the predictor data and specify that the observations are in columns.
X = X'; CVMdl = fitrlinear(X,Y,'ObservationsIn','columns','Lambda',Lambda,... 'Solver','sparsa','Regularization','lasso','CVPartition',cvp); Mdl1 = CVMdl.Trained{1}; numel(Mdl1.Lambda)
ans = 15
Mdl1 is a RegressionLinear model. Because Lambda is a 15-dimensional vector of regularization strengths, you can think of Mdl1 as 15 trained models, one for each regularization strength.
Estimate the test-sample mean squared error for each regularized model.
mse = loss(Mdl1,X(:,idxTest),Y(idxTest),'ObservationsIn','columns');
Higher values of Lambda lead to predictor variable sparsity, which is a good quality of a regression model. Retrain the model using the entire data set and all options used previously, except the data-partition specification. Determine the number of nonzero coefficients per model.
Mdl = fitrlinear(X,Y,'ObservationsIn','columns','Lambda',Lambda,... 'Solver','sparsa','Regularization','lasso'); numNZCoeff = sum(Mdl.Beta~=0);
In the same figure, plot the MSE and frequency of nonzero coefficients for each regularization strength. Plot all variables on the log scale.
figure; [h,hL1,hL2] = plotyy(log10(Lambda),log10(mse),... log10(Lambda),log10(numNZCoeff)); hL1.Marker = 'o'; hL2.Marker = 'o'; ylabel(h(1),'log_{10} MSE') ylabel(h(2),'log_{10} nonzero-coefficient frequency') xlabel('log_{10} Lambda') hold off
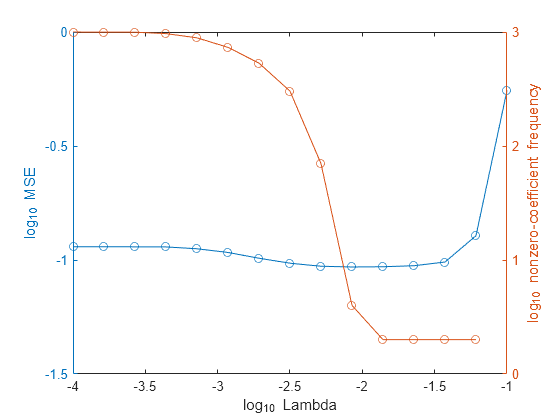
Select the index or indices of Lambda that balance minimal classification error and predictor-variable sparsity (for example, Lambda(11)).
idx = 11; MdlFinal = selectModels(Mdl,idx);
MdlFinal is a trained RegressionLinear model object that uses Lambda(11) as a regularization strength.
Extended Capabilities
The
loss function supports tall arrays with the following usage
notes and limitations:
lossdoes not support talltabledata.
For more information, see Tall Arrays.
This function fully supports GPU arrays. For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2016aloss fully supports GPU arrays.
Starting in R2023b, when you predict or compute the loss, some regression models allow you to specify the predicted response value for observations with missing predictor values. Specify the PredictionForMissingValue name-value argument to use a numeric scalar, the training set median, or the training set mean as the predicted value. When computing the loss, you can also specify to omit observations with missing predictor values.
This table lists the object functions that support the
PredictionForMissingValue name-value argument. By default, the
functions use the training set median as the predicted response value for observations with
missing predictor values.
| Model Type | Model Objects | Object Functions |
|---|---|---|
| Gaussian process regression (GPR) model | RegressionGP, CompactRegressionGP | loss, predict, resubLoss, resubPredict |
RegressionPartitionedGP | kfoldLoss, kfoldPredict | |
| Gaussian kernel regression model | RegressionKernel | loss, predict |
RegressionPartitionedKernel | kfoldLoss, kfoldPredict | |
| Linear regression model | RegressionLinear | loss, predict |
RegressionPartitionedLinear | kfoldLoss, kfoldPredict | |
| Neural network regression model | RegressionNeuralNetwork, CompactRegressionNeuralNetwork | loss, predict, resubLoss, resubPredict |
RegressionPartitionedNeuralNetwork | kfoldLoss, kfoldPredict | |
| Support vector machine (SVM) regression model | RegressionSVM, CompactRegressionSVM | loss, predict, resubLoss, resubPredict |
RegressionPartitionedSVM | kfoldLoss, kfoldPredict |
In previous releases, the regression model loss and predict functions listed above used NaN predicted response values for observations with missing predictor values. The software omitted observations with missing predictor values from the resubstitution ("resub") and cross-validation ("kfold") computations for prediction and loss.
The loss function no longer omits an observation with a
NaN prediction when computing the weighted average regression loss. Therefore,
loss can now return NaN when the predictor data
X or the predictor variables in Tbl
contain any missing values. In most cases, if the test set observations do not contain
missing predictors, the loss function does not return
NaN.
This change improves the automatic selection of a regression model when you use
fitrauto.
Before this change, the software might select a model (expected to best predict the
responses for new data) with few non-NaN predictors.
If loss in your code returns NaN, you can update your code
to avoid this result. Remove or replace the missing values by using rmmissing or fillmissing, respectively.
The following table shows the regression models for which the
loss object function might return NaN. For more details,
see the Compatibility Considerations for each loss
function.
| Model Type | Full or Compact Model Object | loss Object Function |
|---|---|---|
| Gaussian process regression (GPR) model | RegressionGP, CompactRegressionGP | loss |
| Gaussian kernel regression model | RegressionKernel | loss |
| Linear regression model | RegressionLinear | loss |
| Neural network regression model | RegressionNeuralNetwork, CompactRegressionNeuralNetwork | loss |
| Support vector machine (SVM) regression model | RegressionSVM, CompactRegressionSVM | loss |
See Also
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleccione un país/idioma
Seleccione un país/idioma para obtener contenido traducido, si está disponible, y ver eventos y ofertas de productos y servicios locales. Según su ubicación geográfica, recomendamos que seleccione: .
También puede seleccionar uno de estos países/idiomas:
Cómo obtener el mejor rendimiento
Seleccione China (en idioma chino o inglés) para obtener el mejor rendimiento. Los sitios web de otros países no están optimizados para ser accedidos desde su ubicación geográfica.
América
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)